shinymeta — a revolution for reproducibility
Joe Cheng presented shinymeta enabling reproducibility in shiny at useR in July 2019. This is a simple application using shinymeta. You will see how reactivity and reproducibility do not exclude each other. I am really thankful for Joe Cheng realizing the shinymeta project.
Introduction
In 2018 at the R/Pharma conference I first heard of the concept of using quotations. With quotations to make your shiny app code reproducible. This means you can play around in shiny and afterward get the code to generate the exact same outputs as R code. This feature is needed in Pharma. Why is that the case? The pharmaceutical industry needs to report data and analysis to regulatory authorities. I talked about this in several articles already. How great would it be to provide a shiny-app to the regulatory authorities? Great. How great would it be to provide a shiny app that enables them to reproduce every single plot or table? Even better.
Adrian Waddell and Doug Kelkhoff are both my colleges of mine that proposed solutions for this task. Doug built the scriptgloss package which reconstructs static code from shiny apps. Adrian presented a modular shiny-based exploratory framework at R/Pharma 2018. The framework provides dynamic encodings, variable-based filtering, and R-code generation. In this context, I started working out some concepts during my current development project. How to make the code inside a shiny app reproducible? In parallel Doug, Joe Cheng and Carson Sievert worked on a fascinating tool called shinymeta, released on July 11 at the userR conference.
The tool is so fascinating because it created handlers for the task I talked about. It allows changing a simple shiny app into a reproducible shiny app with just a few tweaks. As shiny apps in Pharma have a strong need for this functionality, I am a shiny-developer in Pharma and I wanted to know: How does it work? How good is it?
Let’s create a shiny app relevant in Pharma
As a simple example of a shiny app in Pharma, I will use a linear regression app. The app will detect if a useful linear model can show a correlation between the properties of the patient and the survival rate. Properties of the patient are AGE or GENDER. Survival rates include how long the patient will survive (OS = overall survival), survives without progression (PFS = progression-free survival) or survives without any events occurring (EFS). Each patient can have had all three stages of survival. Let’s create the data sets for this use case with random data:
library(tibble)
library(dplyr)
# Patient listing
pat_data <- list(
SUBJID = 1:200,
STUDYID = c(rep(1, 40), rep(2, 100), rep(3, 60)),
AGE = sample(20:88, 200, replace = T) %>% as.numeric(),
SEX = c(sample(c("M", "F"), 180, replace = T), rep("U", 20)) %>% as.factor()
) %>% as_tibble()
# Days where Overall Survival (OS), Event free survival (EFS) and Progression Free Survival (PFS) happened
event_data <- list(
SUBJID = rep(1:200, 3),
STUDYID = rep(c(rep(1, 40), rep(2, 100), rep(3, 60)), 3),
PARAMCD = c(rep("OS", 200), rep("EFS", 200), rep("PFS", 200)),
AVAL = c(rexp(200, 1 / 100), rexp(200, 1 / 80), rexp(200, 1 / 60)) %>% as.numeric(),
AVALU = rep("DAYS", 600) %>% as.factor()
) %>% as_tibble()
You can see that patient AGE and GENDER (SEX) are randomly distributed. The survival values in days should exponentially decrease. By these distributions, we do not expect to see anything in the data, but this is fine for this example.
 Simple app showing a linear regression of patient data
Simple app showing a linear regression of patient data
Inside the screenshot, you can see the app applied to this data. The app contains the regression plot and the summary of the linear model created with lm . It basically has one input to filter the event_data by PARAMCD. A second input to selects columns from the pat_data . The interesting part of this app is the server function. Inside the server function, there are just two outputs and one reactive value. The reactive performs multiple steps. It generates the formula for the linear model, filters the event_data, selects the pat_data, merges the data sets and calculates the linear model by lm . The two outputs generate a plot and a summary text from the linear model.
# Create a linear model
model_reactive <- reactive({
validate(need(is.character(input$select_regressor), "Cannot work without selected column"))
regressors <- Reduce(function(x, y) call("+", x, y), rlang::syms(input$select_regressor))
formula_value <- rlang::new_formula(rlang::sym("AVAL"), regressors)
event_data_filtered <- event_data %>% dplyr::filter(PARAMCD == input$filter_param)
ads_selected <- pat_data %>% dplyr::select(dplyr::one_of(c(input$select_regressor, c("SUBJID", "STUDYID"))))
anl <- merge(ads_selected, event_data_filtered, by = c("SUBJID", "STUDYID"))
lm(formula = formula_value, data = anl)
})
# Plot Regression vs fitted
output$plot1 <- renderPlot({
plot(model_reactive(), which = 1)
})
# show model summary
output$text1 <- renderPrint({
model_reactive() %>% summary()
})
Of course, you think this app can be easily reproduced by a smart programmer. Now imagine you just see the user-interface and the output. What is missing? Two things are missing:
- How to create the data?
- What is the formula used for creating the linear model?
Let’s make the app reproducible!
By shinymeta and the approach of metaprogramming, we will make the whole app reproducible. Even if shinymeta is still experimental, you will see, right now it works great.
But we need to go step by step. The most important idea behind metaprogramming came from Adrian Waddell. Instead of adding code to your app, you wrap the code in quotations. (Step 1 and the most important).
Creating the data
We can use this for the data added to the app:
data_code <- quote({
# Patient listing
pat_data <- ...
# Days where Overall Survival (OS), Event free survival (EFS) and Progression Free Survival (PFS) happened
event_data <- ...
})
eval(data_code)
Instead of running the code, we wrap it into quote. This will return a call that we can evaluate after by eval . It enables reproducibility. The code that we used to produce the data sets is stored in data_code . We can later on reuse this variable. This variable will allow us to show how the data set was constructed.
Filtering and selecting the data
To enable reproducible filtering and selection we will use the shinymeta functions. Thus we will create a metaReactive returning the merged data set. A metaReactive behaves like a reactive with the difference, that you can get the code used inside back, afterward. This is similar to the principle of quotation. But for the metaReactive you do not need to use an eval function, you can basically stick to the () evaluation, as before.
An important new operator inside the metaReactive is the !! (bang, bang) operator. It allows inserting standard reactive values. It behaves a bit like in the rlang package. You can either use it to inline values from a standard reactive value. Or you can use it to inline metaReactive objects as code. As a summary the operator !! has two functionalities:
- De-reference reactive objects — get their values
- Chain
metaReactive objects by inlining them as code into each other
To get to know the !! operator better, check out the shinymeta vignettes: https://github.com/rstudio/shinymeta/tree/master/vignettes
This code will be used to filter and select and merge the data:
data_set_reactive <- metaReactive({
event_data_filtered <- event_data %>% dplyr::filter(PARAMCD == !!input$filter_param)
ads_selected <- pat_data %>% dplyr::select(dplyr::one_of(c(!!input$select_regressor, c("SUBJID", "STUDYID"))))
merge(ads_selected, event_data_filtered, by = c("SUBJID", "STUDYID"))
})
Inside the code, you can see that the !! operator interacts with the reactive values input$select_regressor and input$filter_param as values. This means we de-reference the reactive value and replace it with its static value. The outcome of this reactive is the merged data set. Of course, this code will not run until we call data_set_reactive() anywhere inside the server function.
Creating the model formula
The formula for the linear model will be created as it was done before:
formula_reactive <- reactive({
validate(need(is.character(input$select_regressor), "Cannot work without selected column"))
regressors <- Reduce(function(x, y) call("+", x, y), rlang::syms(input$select_regressor))
rlang::new_formula(rlang::sym("AVAL"), regressors)
})
It is necessary to check the select regressor value, as without a selection no model can be derived
Creating the linear model
The code to produce the linear model without metaprogramming was as follows:
lm(formula = formula_value, data = anl)
We need to replace formula_value and anl . Additionally replace the reactive with ametaReactive . Therefore we use the function metaReactive2 which allows running standard shiny code before the metaprogramming code. Inside this metaReactive2 it is necessary to check the data and the formula:
validate(need(is.data.frame(data_set_reactive()), "Data Set could not be created"))
validate(need(is.language(formula_reactive()), "Formula could not be created from column selections"))
The metaReactive data_set_reactive can be called like any reactive object. The code to produce the model shall be in meta-programmed because the user wants to see it. The function metaExpr allows this. To get nice reproducible code the call needs to look like this:
metaExpr(bindToReturn = TRUE, {
model_data <- !!data_set_reactive()
lm(formula = !!formula_reactive(), data = model_data)
})
If you do not want to see the whole data set inside the lm call we need to store it inside a variable.
To allow the code to be tracible, you need to put !! in front of the reactive calls. In front of data_set_reactive this allows backtracing the code of data_set_reactive and not only the output value.
Second of all, we can de-reference the formula_reactive by the !! operator. This will directly plug in the formula created into the lm call.
Third, bindToReturn will force shinymeta to write:
var1 <- merge(...)
model_data <- var_1
model_reactive <- lm(formula = AVAL ~ AGE, data = model_data)
instead of
data_set_reactive <- merge(...)
{
model_data <- data_set_reactive
lm(AVAL ~ AGE, data = model_data
}
If you want to read more about the bindToReturn feature, there is an issue on github about the bindToReturn argument. The final model_reactive looks like this:
# Create a linear model
model_reactive <- metaReactive2({
validate(need(is.data.frame(data_set_reactive()), "Data Set could not be created"))
validate(need(is.language(formula_reactive()), "Formula could not be created from column selections"))
metaExpr(bindToReturn = TRUE, {
model_data <- !!data_set_reactive()
lm(formula = !!formula_reactive(), data = model_data)
})
})
Rendering outputs
Last but not least we need to output plots and the text in a reproducible way. Instead of a standard renderPlot and renderPrint function it is necessary to wrap them in metaRender . metaRender enables outputting metaprogramming reactive objects with reproducible code. To get not only the values but also the code of the model, the !! operator is used again.
# Plot Regression vs fitted
output$plot1 <- metaRender(renderPlot, {
plot(!!model_reactive(), which = 1)
})
# show model summary
output$text1 <- metaRender(renderPrint, {
!!model_reactive() %>% summary()
})
Using metaRenderwill make the output a metaprogramming object, too. This allows retrieving the code afterward and makes it reproducible.
Retrieving the code inside the user-interface
IMPORTANT!
Sorry for using capital letters here, but this part is the real part, that makes the app reproducible. By plugging in a “Show R Code” button every user of the app will be allowed to see the code producing outputs. Therefore shinymeta provides the function expandChain . The next section shows how it is used.
In case the user clicks a button, like in this case input$show_r_code a modal with the code should pop up. Inside this modal the expandChain function can handle (1) quoted code and (2)metaRender objects. Each object of such a kind can be used in the … argument of expandChain . It will return a meta-expression. From this meta-expression, the R code used in the app can be extracted. Simply using formatCode() and paste() will make it pretty code show up in the modal.
observeEvent(input$show_r_code, {
showModal(modalDialog(
title = "R Code",
tags$pre(
id = "r_code",
expandChain(
library_code,
data_code,
output$plot1(),
output$text1()
) %>% formatCode() %>% paste(collapse = "\n")
),
footer = tagList(
actionButton("copyRCode", "Copy to Clipboard", `data-clipboard-target` = "#r_code"),
modalButton("Dismiss")
),
size = "l",
easyClose = TRUE
))
})
Please do not forget the () after the metaRender objects.
Final server function and app
 All code can be found at https://github.com/zappingseb/shinymetaTest
All code can be found at https://github.com/zappingseb/shinymetaTest
After going through all steps you can see that the code using shinymeta is not much different from the standard shiny code. Mostly metaReactive , metaReactive2 , metaExpr , metaRender , !! and expandChain are the new functions to learn. Even if the package is still experimental, it does a really good job of making something reactive also reproducible. My favorite functionality is the mixed-use of reactive and metaReactive . By using reactive objects inside meta-code the developer can decide which code goes into the “Show R Code” window and which code runs behind the scenes. You can check yourself by looking into the code of this tutorial. Of course this feature is dangerous, as you might forget to put code in your “Show R Code” window and not all code can be rerun or your reproducible code gets ugly.
The whole code of the tutorial is published on github at: https://github.com/zappingseb/shinymetaTest.
The app runs at https://sebastianwolf.shinyapps.io/shinymetaTest.
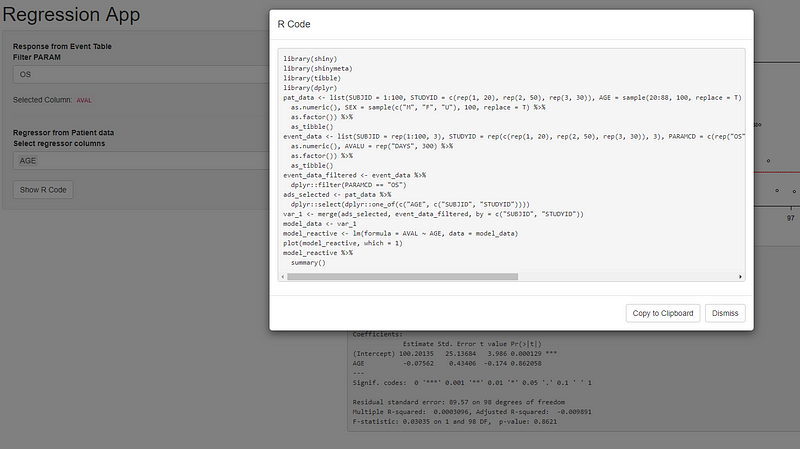 App running at: https://sebastianwolf.shinyapps.io/shinymetaTest/
App running at: https://sebastianwolf.shinyapps.io/shinymetaTest/
Closing words
This was the first time I tried to wrap my own work into a totally new package. The app created inside this example was created within my daily work berfore. The new and experimental package shinymeta allowed switching in ~1 hour from my code to metaprogramming. I did not only switch my implementation, but my implementation also became better due to the package.
Shinymeta will make a huge difference in pharmaceutical shiny applications. One week after the presentation by Joe Cheng I am still impressed by the concept of metaprogramming. And how metaprogramming went into shiny. The package makes shiny really reproducible. It will give guidance for how to use shiny in regulatory fields. Moreover, it will allow more users to code in R, as they can see the code needed for a certain output. Clicking will make them learn.